



@inproceedings{wu2021foodseg,
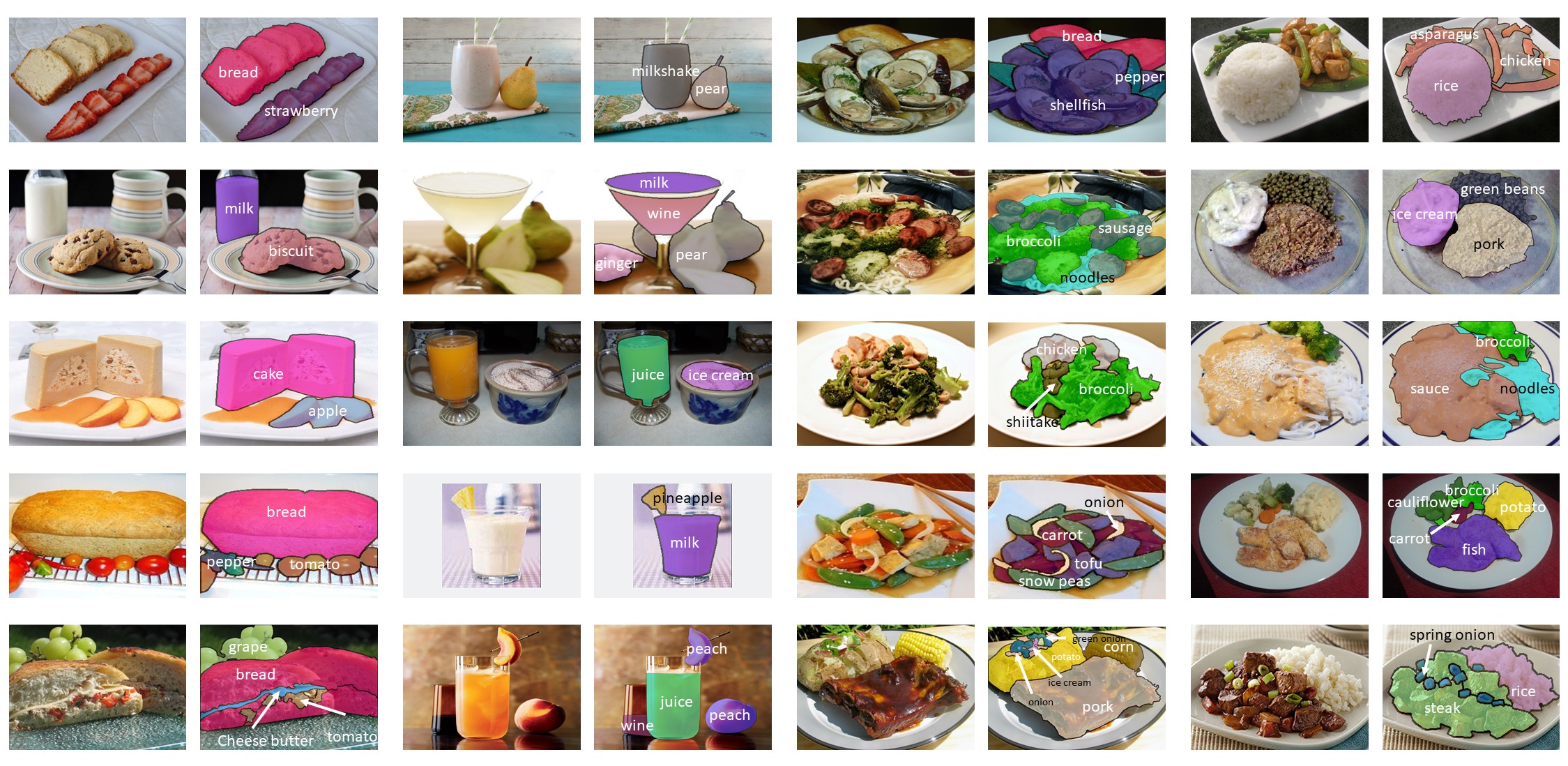
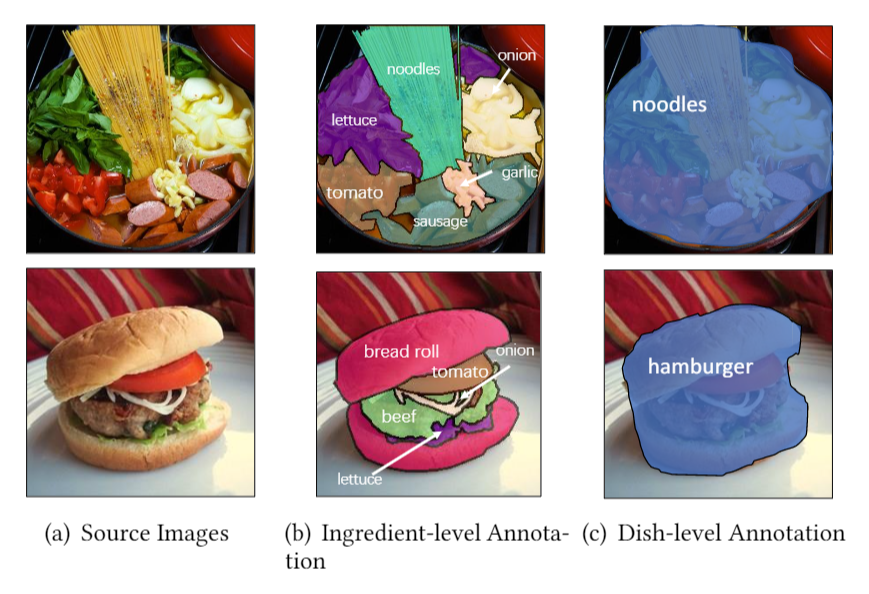
title={A Large-Scale Benchmark for Food Image Segmentation},
author={Wu, Xiongwei and Fu, Xin and Liu, Ying and Lim, Ee-Peng and Hoi, Steven CH and Sun, Qianru},
booktitle={Proceedings of ACM international conference on Multimedia},
year={2021}
}